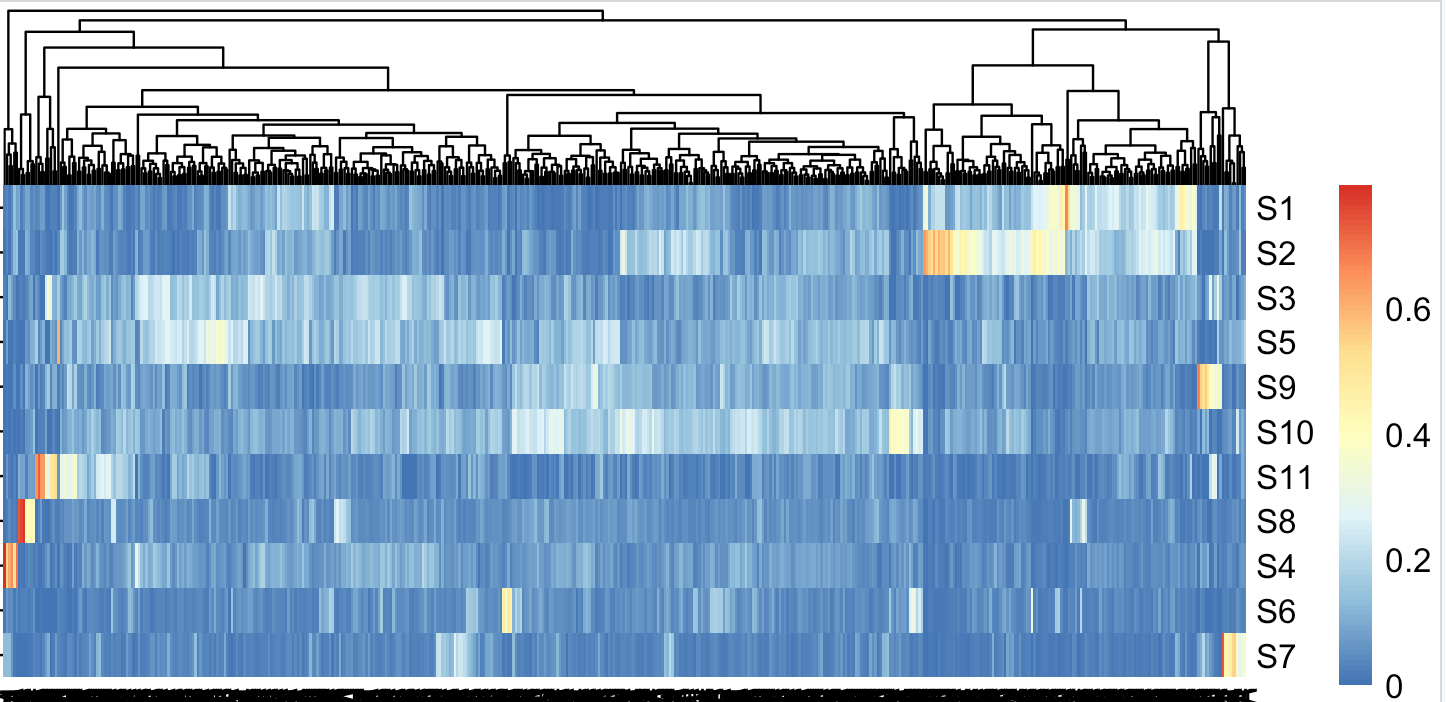
前面我们在教程:使用R包deconstructSigs根据已知的signature进行比例推断,顺利的把508个病人,根据11个signature进行了比例推断,得到的比例矩阵以普通的热图,以及pheatmap包自带的层次聚类如下:
代码是:
rm(list=ls())
options(stringsAsFactors = F)
load(file = 'mut.wt_from_denovo.Rdata')
a2=mut.wt
## 非负矩阵分解识别驱动性signature并绘图
nmf.input <- t(a2)
nmf.input <- nmf.input[setdiff(rownames(nmf.input),"unknown"),] #去掉unknown
# 需要去除那些没有被计算到的signatures
nmf.input <- nmf.input[rowSums(nmf.input)>0,]
library(pheatmap)
pheatmap(nmf.input)
其中代码里面的escc_denovo_results.Rdata文件,来自于教程:使用R包SomaticSignatures进行denovo的signature推断。
可以看到,部分病人的S1,S2比例偏高,很明显,他们应该是一个组,然后剩余的病人可以根据S10进行高低分组,其它signature好像是酱油。
这样的感觉,其实就可以使用NMF算法来实现,尤其是层次聚类并不能很好的把样本进行“泾渭分明”的分组。
第一步:判断最佳分组
就是多次运行 nmf 函数,然后提取cophenetic系数:
library(NMF)
ranks <- 2:10
estim <- lapply(ranks, function(r){
fit <- nmf(nmf.input, r, nrun = 5, seed = 4, method = "lee") # nrun设置为5以免运行时间过长
list(fit = fit, consensus = consensus(fit), .opt = "vp",coph = cophcor(fit))
})
names(estim) <- paste('rank', ranks)
sapply(estim, '[[', 'coph')
就可以绘制随rank变化的cophenetic系数图
png("Cophenetic coefficient for seleting optimal nmf rank.png")
par(cex.axis=1.5)
plot(ranks, sapply(estim, '[[', 'coph'),
xlab="", ylab="", type="b",
col="red", lwd=4,xaxt="n")
axis(side = 1, at=1:10)
title(xlab="number of clusters", ylab="Cophenetic coefficient", cex.lab=1.5)
dev.off()
如下所示:
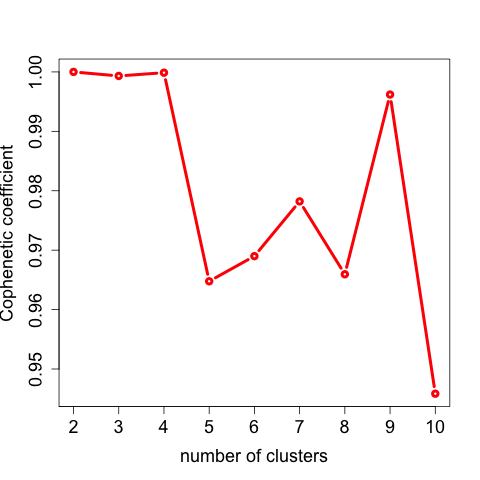
理论上,根据cophenetic得分选择rank=4,因为它是拐点,但是0元,10小时教学视频直播《跟着百度李彦宏学习肿瘤基因组测序数据分析》 这个文献里面设置为3。这里,我们就尊重文献结果吧。
第二步:筛选signature
前面我们的508个病人,都是11个signature,但是呢,我们的NMF算法运行过后,可以看到有一些signature其实对样本聚类分组并没有意义,所以我们需要挑选一下,代码如下:
rank <- 3
seed <- 2019620
rownames(nmf.input) <- gsub("Signature","Sig",rownames(nmf.input)) # 行名简化
mut.nmf <- nmf(nmf.input,
rank = rank,
seed = seed,
method = "lee")
index <- extractFeatures(mut.nmf,"max")
sig.order <- unlist(index)
得到的结果如下;
> index
[[1]]
[1] 2 1
[[2]]
[1] 5 3 11 4
[[3]]
[1] 10 9
attr(,"method")
[1] "max"
> sig.order
[1] 2 1 5 3 11 4 10 9
可以看到,nmf算法得到的3类,都是有各自偏重的signature,11个signature也只有8个需要后续进行分析。如下图,可以看到不同nmf类有各自的偏重的signature。
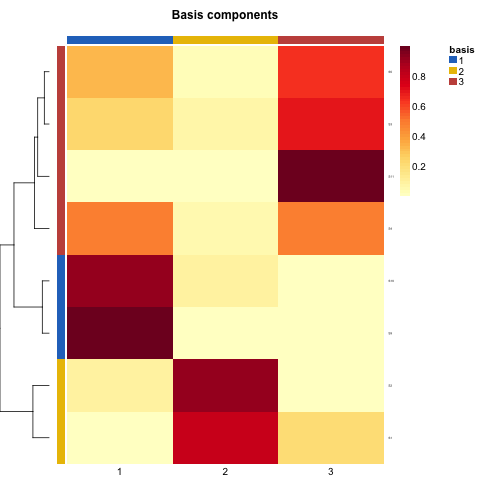
第三步:使用挑选出的signature再次NMF
nmf.input2 <- nmf.input[sig.order,]
library(pheatmap)
pheatmap(nmf.input2,cluster_rows = T,cluster_cols = F)
mut.nmf2 <- nmf(nmf.input2,
rank = rank,
seed = seed,
method = "lee")
group <- predict(mut.nmf2) # 提出亚型
table(group)
这个时候,病人的分类就是最后的nmf区分成为的3类。泾渭分明,如下:
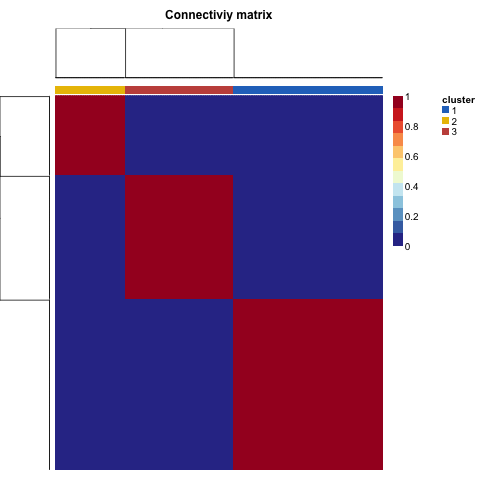
番外:一些可视化函数
主要是继续参考每个nmf类里面的不同signature的比例,以及不同nmf类的相关性热图。
sample.order <- names(group[order(group)])
#设置颜色
jco <- c("#2874C5","#EABF00","#C6524A","#868686")
png(file = "consensusmap.png")
consensusmap(mut.nmf2,
labRow = NA,
labCol = NA,
annCol = data.frame("cluster"=group[colnames(nmf.input)]),
annColors = list(cluster=c("1"=jco[1],"2"=jco[2],"3"=jco[3],"4"=jco[4])))
dev.off()
png(file = "basismap.png" )
# 从这张图可以比较清晰地看到各亚型中的驱动signature(深色),与下面的nmf heatmap对应
basismap(mut.nmf2,
cexCol = 1,
cexRow = 0.3,
annColors=list(c("1"=jco[1],"2"=jco[2],"3"=jco[3],"4"=jco[4])))
dev.off()
aheatmap(as.matrix(nmf.input2[,sample.order]),
Rowv=NA,
Colv=NA,
annCol = data.frame("cluster"=group[sample.order]),
annColors = list(cluster=c("1"=jco[1],"2"=jco[2],"3"=jco[3],"4"=jco[4])),
color=c("#EAF0FA","#6081C3","#3454A7"), # 例文的蓝色渐变
revC=TRUE,
cexCol = 0.3,
cexRow = 0.3,
filename = "NMF_heatmap.pdf")
ac=data.frame(group=paste0('NMF_',group))
rownames(ac)=colnames(nmf.input2)
pheatmap(nmf.input2,show_colnames = F,annotation_col = ac)
pheatmap(nmf.input2[,sample.order],
show_colnames = F,cluster_cols = F,annotation_col = ac)
pheatmap(nmf.input2,show_colnames = F,annotation_col = ac,
filename = "NMF_heatmap1.pdf")
pheatmap(nmf.input2[,sample.order],
show_colnames = F,cluster_cols = F,annotation_col = ac,
filename = "NMF_heatmap2.pdf")
dev.off()
可视化是一条永无止境的路。大家加油哈!